Software-Supported USER Cloning Strategies for Site-Directed Mutagenesis and DNA Assembly | ACS Synthetic Biology

Development of a sensitive, quantitative assay with broad subtype specificity for detection of total HIV-1 nucleic acids in plasma and PBMC | Scientific Reports

IJMS | Free Full-Text | The T850D Phosphomimetic Mutation in the Androgen Receptor Ligand Binding Domain Enhances Recruitment at Activation Function 2

Recognition motif and mechanism of ripening inhibitory peptides in plant hormone receptor ETR1 | Scientific Reports

IJMS | Free Full-Text | Secondary Structures of the Transmembrane Domain of SARS-CoV-2 Spike Protein in Detergent Micelles

Enhancing the Thermal Robustness of an Enzyme by Directed Evolution: Least Favorable Starting Points and Inferior Mutants Can Map Superior Evolutionary Pathways - Gumulya - 2011 - ChemBioChem - Wiley Online Library

Half-life (t 1/2 ) Calculation from Thermal Inactivation Kinetics of... | Download Scientific Diagram

Microorganisms | Free Full-Text | Critical Role of 3′-Downstream Region of pmrB in Polymyxin Resistance in Escherichia coli BL21(DE3)
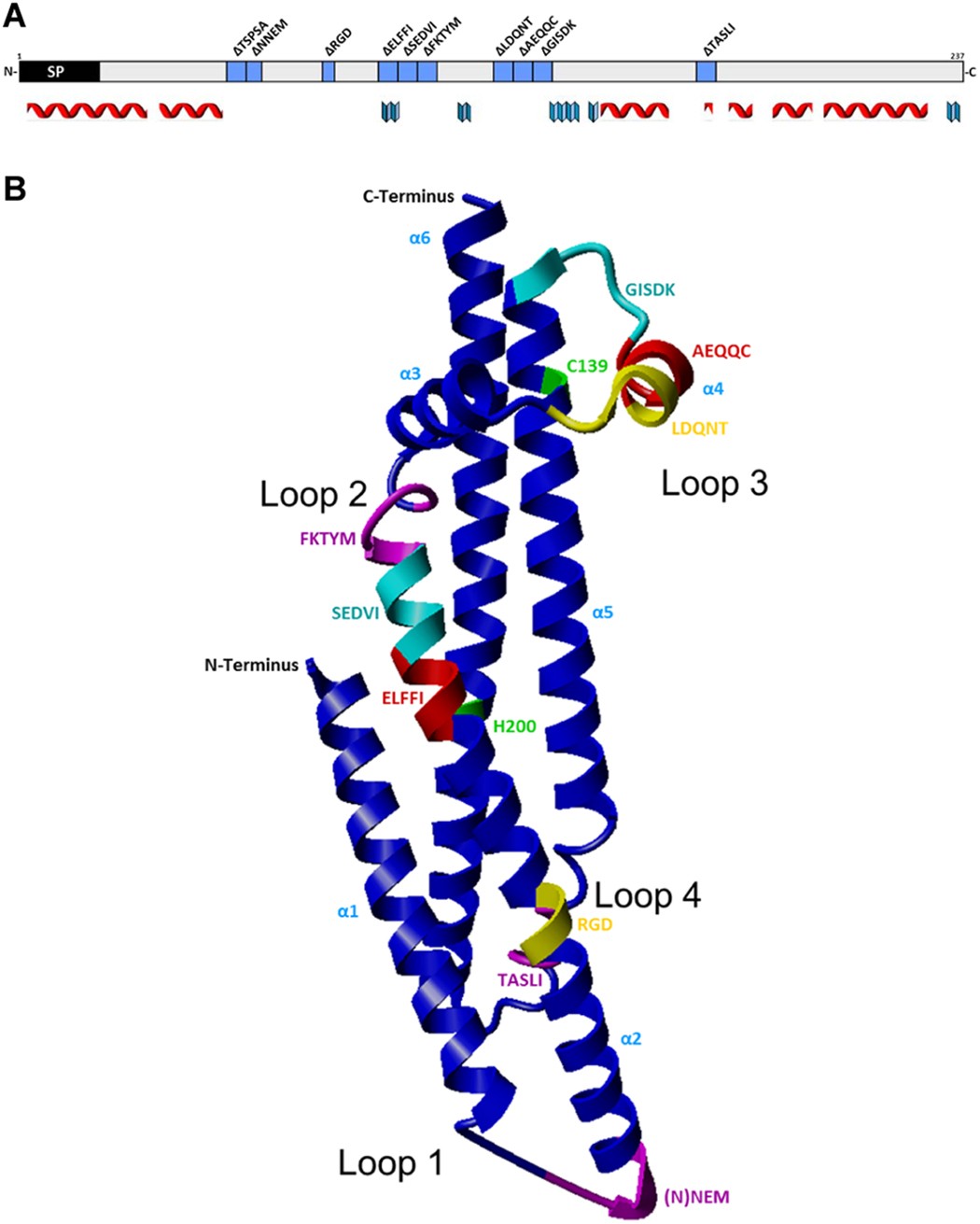
Systematic site-directed mutagenesis of the Helicobacter pylori CagL protein of the Cag type IV secretion system identifies novel functional domains | Scientific Reports

Software-Supported USER Cloning Strategies for Site-Directed Mutagenesis and DNA Assembly | ACS Synthetic Biology

Optimization of Furin Inhibitors To Protect against the Activation of Influenza Hemagglutinin H5 and Shiga Toxin | Journal of Medicinal Chemistry

Photocrosslinking-induced CRAC channel-like Orai1 activation independent of STIM1 | Nature Communications